Deciphering the Cell Code that perpetuates animal life
The information required to perpetuate animal life must be contained even within a single cell that begins each generation (e.g. fertilized egg in sexually reproducing animals). The DNA within this cell encodes what can be made but does not determine what is made. What is made depends on how the cell interprets the DNA sequence, and requires instructions and machinery in addition to the DNA. Thus, life 'begins' with two distinct forms of information: the linear DNA sequence that is faithfully replicated during cell divisions, and a three-dimensional arrangement of molecules that dictates what is made using DNA and changes during development but returns to a similar configuration at the start of each generation. These two interdependent stores of information – one replicating with every cell division and the other cycling with a period of one generation – coevolve and together can be thought of as forming a cell code for making an organism.
Understanding the cell code and how it is propagated during development such that it is recreated in a similar configuration at the start of each generation has implications for evolution, origins of inherited diseases, and consequences of genome engineering. Taking advantage of our recent ability to induce transgenerational epigenetic inheritance, i.e. modify non-genetic aspects of the cell code, our goal is to use reductionist, systems, and engineering approaches to understand the information required to build and perpetuate an animal.
Hints that we can change non-genetic aspects of the cell code come from phenomena like paramutation and RNA interference, where changes in gene regulatory information can persist across generations. Both phenomena rely on RNA for sequence specificity and the details of how they affect gene expression are still being worked out. But, this ability to pass non-genetic information across generational boundaries and recent advances in single cell analyses, genome editing, and epigenome engineering enable us to begin deciphering the cell code that perpetuates life.
Our research using the simple worm C. elegans in the following four areas provides tools and a framework that guides our approach to this central problem: Transgenerational RNA Silencing, Extracellular RNA, Tiny RNAs, and Tissue Homogeneity.
Overview: We found that double-stranded RNA (dsRNA) expressed in neurons can enter the germline and silence a gene of matching sequence and that the silencing can last for more than 25 generations (Devanapally et al., 2015). The dsRNA exported from neurons in an animal, injected into an animal, or ingested by an animal are all expected to reach the body cavity that surrounds most tissues in C. elegans. Analysis of genetic requirements for silencing by ingested dsRNA revealed that each somatic cell needs long dsRNA for silencing (Raman et al., 2017). Yet, we found that any extracellular material including RNA can directly reach progeny after entry into oocytes (Marré et al., 2016), suggesting that dsRNA made in neurons could similarly enter oocytes, reach progeny, and then initiate transgenerational gene silencing. The multi-generational maintenance of gene silencing requires a nuclear Argonaute protein (Devanapally et al., 2015) that binds antisense small RNAs (~22 nucleotides), which are likely made in every generation.
Quantifying small RNAs associated with silencing is possible through next-generation RNA sequencing (RNA-seq) but the analysis of the resultant data can be challenging. We developed bioinformatic approaches for the clear analysis of RNA-seq data and discovered a new class of small RNAs that are shorter than 18 nucleotides (Blumenfeld & Jose, 2016). Experimental and theoretical considerations suggest that there are additional such tiny RNAs and we have improved northern blotting to enable their independent analysis (Choi et al., 2017).
Even if the information to express or silence a gene is passed from one generation to the next, in every generation, animals face the challenge of keeping the expression of a gene similar in cells within a tissue despite cell divisions. We found that a variable set of cells become susceptible to silencing of repetitive DNA in the absence of factors that inhibit RNA silencing and that the dsRNAs that trigger such silencing can move between cells in the embryo (Le et al., 2016).
Thus, we have begun to learn how to use RNA in one generation to affect the expression of a matching gene in subsequent generations and have developed tools to effectively analyze small RNAs associated with gene silencing in every generation.
Building on these past studies, we plan to change the expression of a specific gene without affecting its sequence and discover the factors that restrict or allow the persistence of such changes for multiple generations. As a complementary approach, we plan to use CRISPR-based genome editing to alter DNA to discover the sequence characteristics that restrict or allow transgenerational reprogramming.
Transgenerational RNA Silencing
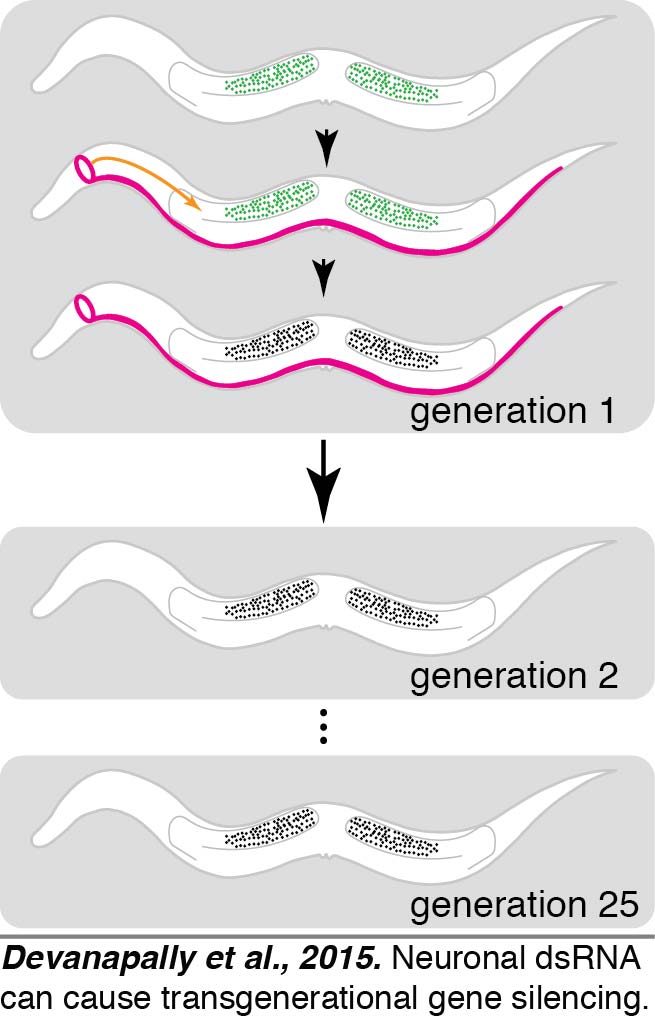 What happens to our bodies is typically thought to not influence the biology of our descendants. The reason for this view is that the somatic cells that generate our body are separated early in development from the germ cells that make sperm or egg. This separation had led to the belief in a barrier to communication between somatic cells and germ cells, a concept known as the Weismann Barrier. Our work establishes a way for somatic cells to communicate gene-specific information to germ cells and breaks the Weismann Barrier (Devanapally et al., 2015).
What happens to our bodies is typically thought to not influence the biology of our descendants. The reason for this view is that the somatic cells that generate our body are separated early in development from the germ cells that make sperm or egg. This separation had led to the belief in a barrier to communication between somatic cells and germ cells, a concept known as the Weismann Barrier. Our work establishes a way for somatic cells to communicate gene-specific information to germ cells and breaks the Weismann Barrier (Devanapally et al., 2015).
We showed that neurons (magenta) of the worm C. elegans can send gene-specific messages in the form of dsRNA to the germline (green) and cause transgenerational gene silencing that can last for more than 25 generations. Neuronal dsRNAs enter germ cells through an importer protein called SID-1, which has homologs in most animals, including humans. These results can have profound implications for our understanding of evolution and behavior. We have begun a UMD-funded program called the Transgenerational Brain Initiative that trains ~35 undergraduates each year in original research aimed at discovering which neuron(s) are the best exporters of dsRNA to the germline.
Current models for the transgenerational inheritance of silencing in C. elegans propose chromatin modifications associated with the production of small RNAs that reinforce these modifications in each generation. Most aspects of this model, however, remain to be tested and only a few protein factors required for this process are known. RNAs transcribed from the C. elegans genome that move from neurons to the germline to cause transgenerational gene silencing are unknown.
Extracellular RNA
Cells can communicate with each other through the direct transport of macromolecules (e.g. RNA) in many organisms, including humans. Secreted vesicles with such macromolecules that are found in blood are being hotly pursued as valuable diagnostic markers for the health of internal organs. However, the mechanisms that enable this novel mode of cell communication are not well understood (see Jose, 2015 for review). The transport of RNA between cells had been inferred in C. elegans since the discovery of RNA interference. While such systemic silencing was apparent when dsRNA was fed to worms or injected into them, it was unclear whether dsRNA expressed in one cell can typically cause silencing in another cell. We established that many somatic cell types can export RNAs that enter other cell types and cause gene silencing (Jose et al., 2009) and identified a conserved tyrosine kinase as a regulator of dsRNA import into cells (Jose et al., 2012).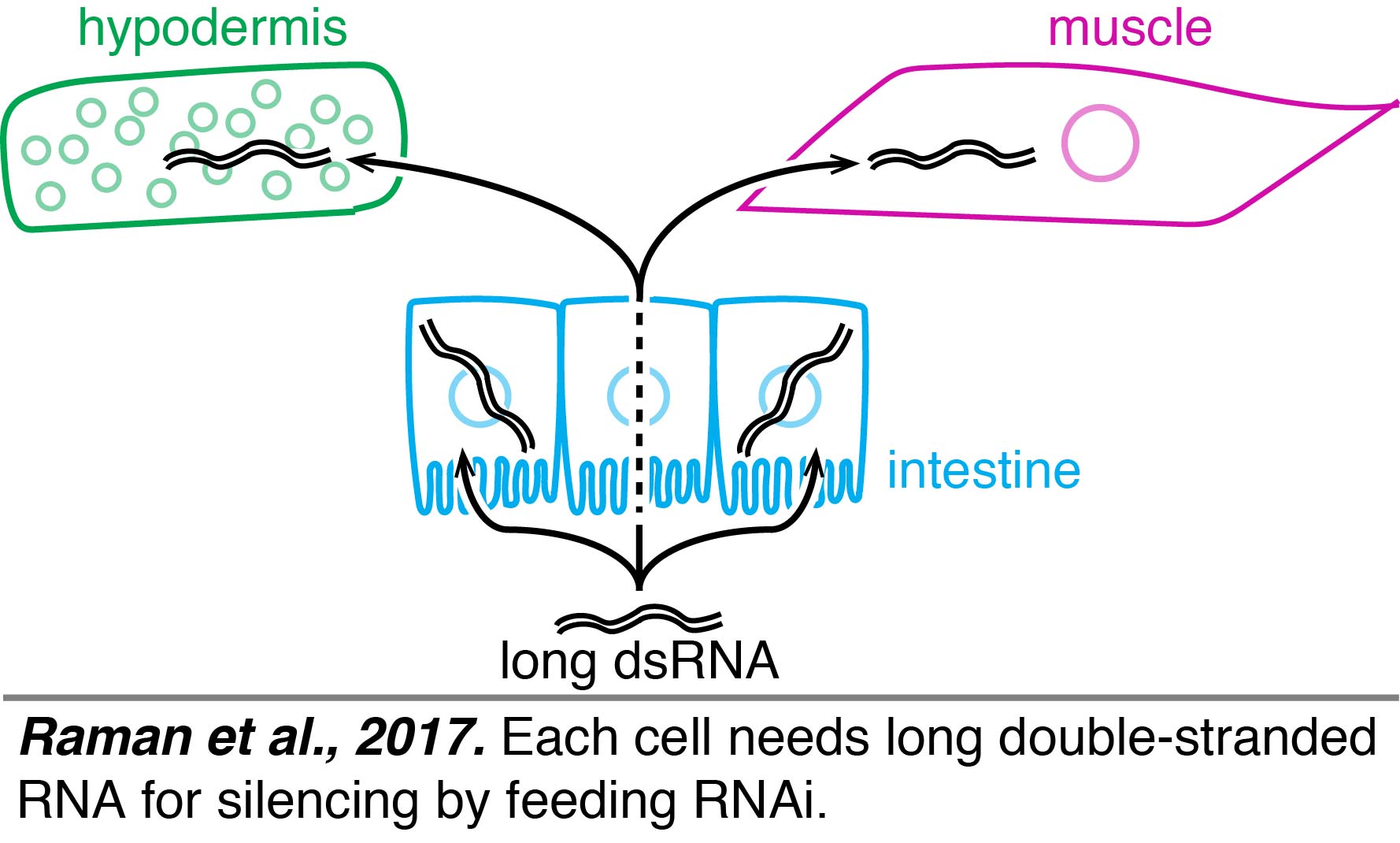
Tissue-specific rescue experiments had suggested that there were two forms of dsRNA that could move between cells – long dsRNA and short dsRNA – when long dsRNA was expressed from neurons (Jose et al., 2011). A similar conclusion was reached when ingested dsRNA was used to initiate silencing, suggesting that long dsRNA could enter a cell, be processed into short dsRNAs, and subsequently the short dsRNAs could move to other cells. This "transit" model for silencing by ingested dsRNA relied entirely on experiments that used repetitive transgenes for tissue-specific rescue, which can result in misexpression even when well-characterized promoters are used because of new promoters that can arise from DNA rearrangements (Le et al., 2016). Furthermore, expression from any repetitive transgene within a tissue can inhibit silencing of some genes by ingested dsRNA within that tissue (Raman et al., 2017), which could complicate interpretations of many past experiments that used feeding RNAi to study diverse problems in biology. A re-evaluation of the tissue-specific rescue experiments using single-copy transgenes showed that each cell needs long dsRNA for silencing by feeding RNAi (Raman et al., 2017).
Regardless of how dsRNAs are delivered into the worm – expression within a tissue, injection, 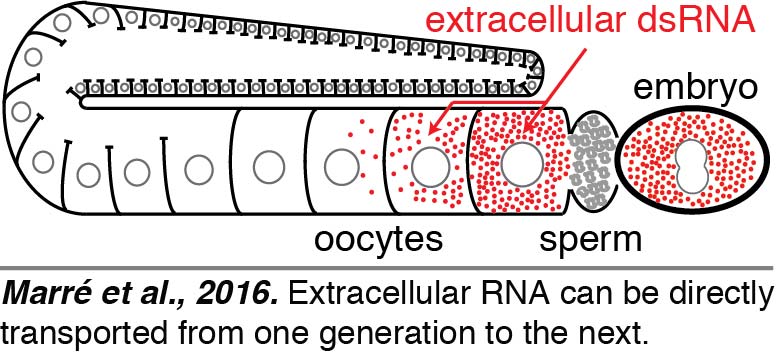 ingestion, or soaking – dsRNAs are expected to reach the body cavity that surrounds all tissues in C. elegans. To directly visualize the fate of such extracellular dsRNAs, we injected fluorescently labeled 50-bp dsRNA into the body cavity. We found that dsRNA entered oocytes along with yolk and reached progeny (Marré et al., 2016). Surprisingly, we found that such delivery of extracellular RNA from parent to progeny did not require SID-1-mediated entry into the cytosol within the parent. Thus, the dsRNA that enters oocytes along with yolk can presumably be held within intracellular vesicles and reach embryos. These results raise the intriguing possibility that extracellular RNAs that are secreted – potentially in response to changes in a parent – can directly reach progeny and regulate gene expression.
ingestion, or soaking – dsRNAs are expected to reach the body cavity that surrounds all tissues in C. elegans. To directly visualize the fate of such extracellular dsRNAs, we injected fluorescently labeled 50-bp dsRNA into the body cavity. We found that dsRNA entered oocytes along with yolk and reached progeny (Marré et al., 2016). Surprisingly, we found that such delivery of extracellular RNA from parent to progeny did not require SID-1-mediated entry into the cytosol within the parent. Thus, the dsRNA that enters oocytes along with yolk can presumably be held within intracellular vesicles and reach embryos. These results raise the intriguing possibility that extracellular RNAs that are secreted – potentially in response to changes in a parent – can directly reach progeny and regulate gene expression.
How dsRNA is exported from cells and the basis for differences in gene silencing by long and short dsRNAs are unknown. RNAs transcribed from the C. elegans genome or from ingested bacteria that accumulate extracellularly or that reach progeny are also unknown.
Tiny RNA
Small RNAs are major regulators of gene expression in C. elegans, and there are millions of them. They are typically identified using RNA-seq, but the effective analysis of the resulting data is a challenge. Therefore, we created a set of programs called PACER (Programs for Analysis of C. elegans small RNAs) to effectively analyze small RNAs detected through RNA-seq (Blumenfeld & Jose, 2016). Using PACER, we gained insight 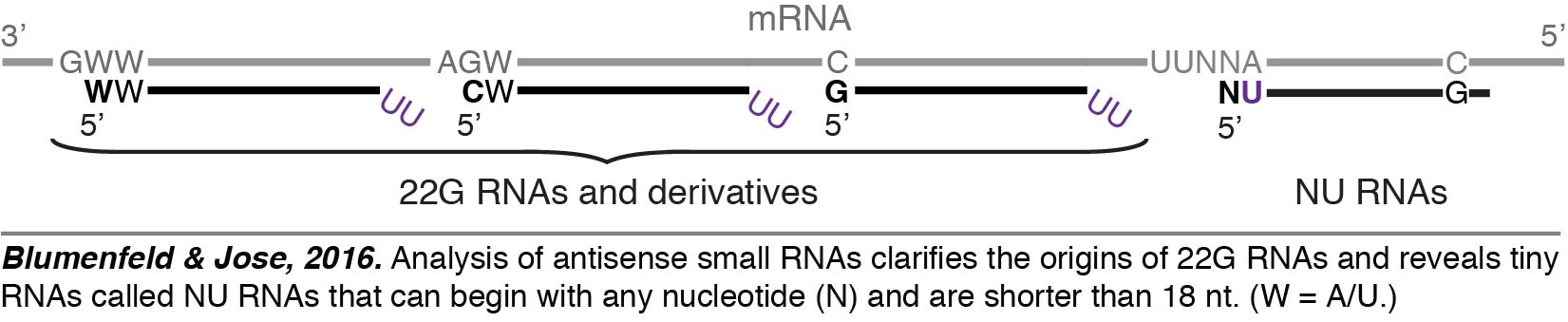 into the origins of the diverse set of antisense small RNAs called 22G RNAs that are used for sequence-specific gene regulation in C. elegans and discovered a new class of tiny RNAs that are shorter than 18 nucleotides (18-nt) called NU RNAs (pronounced “new RNAs”). The minimum length of a small RNA that is required for sequence-specific regulation depends on the sequence space that needs to be searched. Thus, we propose that RNAs shorter than 18-nt could be used for sequence-specific regulation – particularly in organisms with small genomes – and represent an underexplored world of functional tiny RNAs. This speculation is supported by our detection of NU RNAs despite most RNA-seq experiments selecting against RNAs that are less than 18-nt.
into the origins of the diverse set of antisense small RNAs called 22G RNAs that are used for sequence-specific gene regulation in C. elegans and discovered a new class of tiny RNAs that are shorter than 18 nucleotides (18-nt) called NU RNAs (pronounced “new RNAs”). The minimum length of a small RNA that is required for sequence-specific regulation depends on the sequence space that needs to be searched. Thus, we propose that RNAs shorter than 18-nt could be used for sequence-specific regulation – particularly in organisms with small genomes – and represent an underexplored world of functional tiny RNAs. This speculation is supported by our detection of NU RNAs despite most RNA-seq experiments selecting against RNAs that are less than 18-nt.
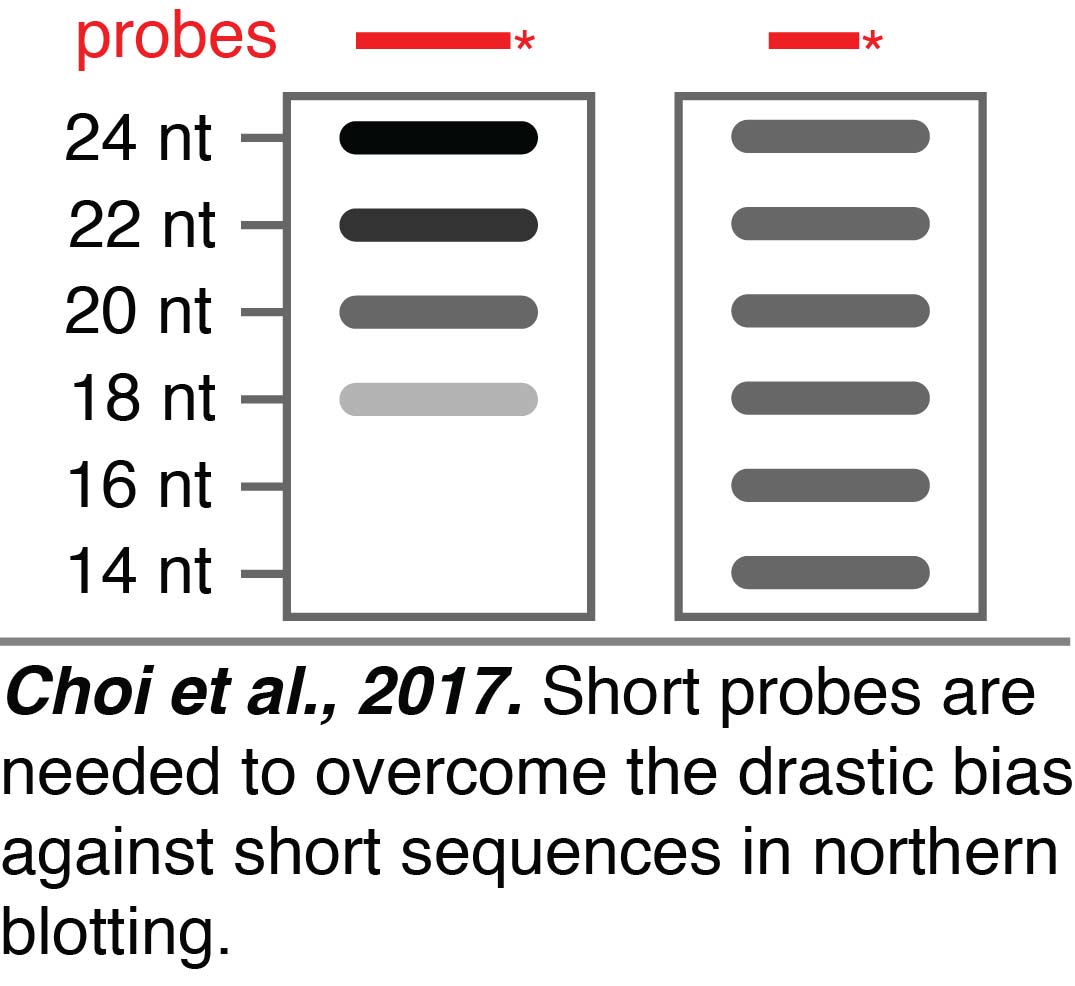 When analyzing RNA-seq data, we discovered that some of the sequences present in a dataset could be generated during the preparatory steps required for RNA-seq, highlighting the need for additional approaches to study small RNAs. Small RNAs can be independently examined using northern blotting. We found that northern blotting of small RNAs can have a drastic bias against short sequences (Choi et al., 2017). We greatly reduced the bias from ~360 fold against 18 nt RNAs compared to 24 nt RNAs when using a 24 nt probe to a maximum of ~4 fold across the entire range of 24 nt to 14 nt RNAs by using 15 nt probes and low hybridization temperatures. This improved northern blotting will be useful for the independent evaluation of any tiny RNAs that we discover when we perform RNA-seq without selecting against RNAs that are less than 18-nt.
When analyzing RNA-seq data, we discovered that some of the sequences present in a dataset could be generated during the preparatory steps required for RNA-seq, highlighting the need for additional approaches to study small RNAs. Small RNAs can be independently examined using northern blotting. We found that northern blotting of small RNAs can have a drastic bias against short sequences (Choi et al., 2017). We greatly reduced the bias from ~360 fold against 18 nt RNAs compared to 24 nt RNAs when using a 24 nt probe to a maximum of ~4 fold across the entire range of 24 nt to 14 nt RNAs by using 15 nt probes and low hybridization temperatures. This improved northern blotting will be useful for the independent evaluation of any tiny RNAs that we discover when we perform RNA-seq without selecting against RNAs that are less than 18-nt.
The evolutionarily selected functions of NU RNAs, if any, are unknown. Whether there are additional such tiny RNAs in C. elegans and whether they are found in mammals and other organisms with larger genomes are also unknown.
Tissue Homogeneity
Through our studies on RNA silencing of repetitive DNA, we have stumbled upon a fundamental question that has received little attention: given the numerous components of a cell, how do animals keep any two cells similar within a tissue? We discovered that inhibitors of RNA silencing that are inherited from parents as well as made in developing progeny act in the embryo to eliminate variation between cells.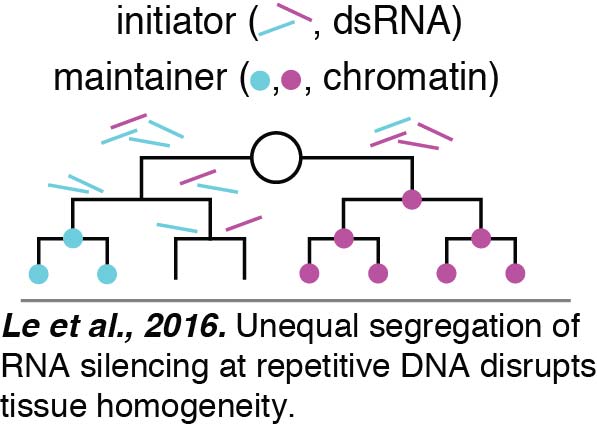 In the absence of such factors, an initiator of gene silencing, likely dsRNA, is segregated unequally between cells during proliferative cell divisions required to make a tissue. This initiator causes threshold-dependent gene silencing. Consistent with the initiator being dsRNA, fewer cells show silencing when SID-1 is removed to block the spread of dsRNA between cells. Once initiated, the silencing is maintained by chromatin-mediated mechanisms despite DNA replication and cell division (Le et al., 2016).
In the absence of such factors, an initiator of gene silencing, likely dsRNA, is segregated unequally between cells during proliferative cell divisions required to make a tissue. This initiator causes threshold-dependent gene silencing. Consistent with the initiator being dsRNA, fewer cells show silencing when SID-1 is removed to block the spread of dsRNA between cells. Once initiated, the silencing is maintained by chromatin-mediated mechanisms despite DNA replication and cell division (Le et al., 2016).
Once different tissues have been specified through unequal cell divisions and signaling in the early embryo, subsequent cell divisions within a tissue could have to generate similar cells. Given the fundamental nature of this requirement, we suspect that many more processes in cells are under similar control to ensure a clean switch from differentiation to epigenetic memory during proliferative cell divisions that generate a tissue. Loss of such developmental mechanisms could potentially cause diseases later in life because of the generation of a variable population of cells within a tissue. This consideration is particularly relevant for age-related disease such as cancer that strike single cells apparently at random.
Understanding how such developmental mechanisms maintain the expression state of a gene will be essential to control the precise expression or silencing of a gene in every generation.
Back to research
Last updated: Aug 2017